ARG53328
anti-MUC1 / EMA antibody
anti-MUC1 / EMA antibody for ICC/IF,IHC-Formalin-fixed paraffin-embedded sections,Immunoprecipitation and Human,Mouse
Cancer antibody; Controls and Markers antibody; Signaling Transduction antibody; Epithelial Marker antibody
Overview
| Product Description | Rabbit Polyclonal antibody recognizes MUC1 / EMA |
|---|---|
| Tested Reactivity | Hu, Ms |
| Tested Application | ICC/IF, IHC-P, IP |
| Host | Rabbit |
| Clonality | Polyclonal |
| Isotype | IgG |
| Target Name | MUC1 / EMA |
| Antigen Species | Human |
| Immunogen | Synthetic peptide from the cytoplasmic tail of Human MUC1 / EMA. |
| Conjugation | Un-conjugated |
| Alternate Names | MUC1-NT; EMA; Mucin-1; MAM6; PEM; Peanut-reactive urinary mucin; CD227; MUC-1/SEC; Breast carcinoma-associated antigen DF3; MUC-1/X; Cancer antigen 15-3; H23AG; CD antigen CD227; MCKD; Carcinoma-associated mucin; Polymorphic epithelial mucin; MUC-1; MUC1-alpha; KL-6; Tumor-associated epithelial membrane antigen; MUC1-CT; ADMCKD1; Episialin; PUM; Tumor-associated mucin; PEMT; MCKD1; ADMCKD; MCD; Krebs von den Lungen-6; CA 15-3; MUC1-beta; MUC1/ZD |
Application Instructions
| Application Suggestion |
|
||||||||
|---|---|---|---|---|---|---|---|---|---|
| Application Note | IHC-P: Incubation Time: 10 min at RT. * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. |
||||||||
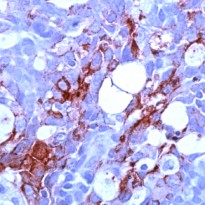
| Positive Control | Breast Carcinoma |
Properties
| Form | Liquid |
|---|---|
| Purification | Immunogen affinity purified |
| Buffer | PBS (pH 7.6), 1% BSA and <0.1% Sodium azide |
| Preservative | <0.1% Sodium azide |
| Stabilizer | 1% BSA |
| Storage Instruction | For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C or below. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. |
| Note | For laboratory research only, not for drug, diagnostic or other use. |
Bioinformation
| Database Links | |
|---|---|
| Gene Symbol | MUC1 |
| Gene Full Name | mucin 1, cell surface associated |
| Background | This gene encodes a membrane-bound protein that is a member of the mucin family. Mucins are O-glycosylated proteins that play an essential role in forming protective mucous barriers on epithelial surfaces. These proteins also play a role in intracellular signaling. This protein is expressed on the apical surface of epithelial cells that line the mucosal surfaces of many different tissues including lung, breast stomach and pancreas. This protein is proteolytically cleaved into alpha and beta subunits that form a heterodimeric complex. The N-terminal alpha subunit functions in cell-adhesion and the C-terminal beta subunit is involved in cell signaling. Overexpression, aberrant intracellular localization, and changes in glycosylation of this protein have been associated with carcinomas. This gene is known to contain a highly polymorphic variable number tandem repeats (VNTR) domain. Alternate splicing results in multiple transcript variants.[provided by RefSeq, Feb 2011] |
| Function | The alpha subunit has cell adhesive properties. Can act both as an adhesion and an anti-adhesion protein. May provide a protective layer on epithelial cells against bacterial and enzyme attack. The beta subunit contains a C-terminal domain which is involved in cell signaling, through phosphorylations and protein-protein interactions. Modulates signaling in ERK, SRC and NF-kappa-B pathways. In activated T-cells, influences directly or indirectly the Ras/MAPK pathway. Promotes tumor progression. Regulates TP53-mediated transcription and determines cell fate in the genotoxic stress response. Binds, together with KLF4, the PE21 promoter element of TP53 and represses TP53 activity. [UniProt] |
| Cellular Localization | Cytoplasmic and cell surface. [UniProt] |
| Highlight | Related Antibody Duos and Panels: ARG30305 Epithelial Marker Antibody Duo (Muc-1, EpCAM) Related products: MUC1 antibodies; MUC1 ELISA Kits; MUC1 Duos / Panels; Anti-Rabbit IgG secondary antibodies; Related news: Lymphoma |
| Research Area | Cancer antibody; Controls and Markers antibody; Signaling Transduction antibody; Epithelial Marker antibody |
| Calculated MW | 122 kDa |
| PTM | Highly glycosylated (N- and O-linked carbohydrates and sialic acid). O-glycosylated to a varying degree on serine and threonine residues within each tandem repeat, ranging from mono- to penta-glycosylation. The average density ranges from about 50% in human milk to over 90% in T47D breast cancer cells. Further sialylation occurs during recycling. Membrane-shed glycoproteins from kidney and breast cancer cells have preferentially sialyated core 1 structures, while secreted forms from the same tissues display mainly core 2 structures. The O-glycosylated content is overlapping in both these tissues with terminal fucose and galactose, 2- and 3-linked galactose, 3- and 3,6-linked GalNAc-ol and 4-linked GlcNAc predominating. Differentially O-glycosylated in breast carcinomas with 3,4-linked GlcNAc. N-glycosylation consists of high-mannose, acidic complex-type and hybrid glycans in the secreted form MUC1/SEC, and neutral complex-type in the transmembrane form, MUC1/TM. Proteolytic cleavage in the SEA domain occurs in the endoplasmic reticulum by an autoproteolytic mechanism and requires the full-length SEA domain as well as requiring a Ser, Thr or Cys residue at the P + 1 site. Cleavage at this site also occurs on isoform MUC1/X but not on isoform MUC1/Y. Ectodomain shedding is mediated by ADAM17. Dual palmitoylation on cysteine residues in the CQC motif is required for recycling from endosomes back to the plasma membrane. Phosphorylated on tyrosines and serine residues in the C-terminal. Phosphorylation on tyrosines in the C-terminal increases the nuclear location of MUC1 and beta-catenin. Phosphorylation by PKC delta induces binding of MUC1 to beta-catenin/CTNNB1 and thus decreases the formation of the beta-catenin/E-cadherin complex. Src-mediated phosphorylation inhibits interaction with GSK3B. Src- and EGFR-mediated phosphorylation on Tyr-1229 increases binding to beta-catenin/CTNNB1. GSK3B-mediated phosphorylation on Ser-1227 decreases this interaction but restores the formation of the beta-cadherin/E-cadherin complex. On T-cell receptor activation, phosphorylated by LCK. PDGFR-mediated phosphorylation increases nuclear colocalization of MUC1CT and CTNNB1. The N-terminal sequence has been shown to begin at position 24 or 28. |
Images (1) Click the Picture to Zoom In